DECONSTRUCTING EVOLUTION THROUGH THE POPULATION PALAEOGENOMIC WINDOW
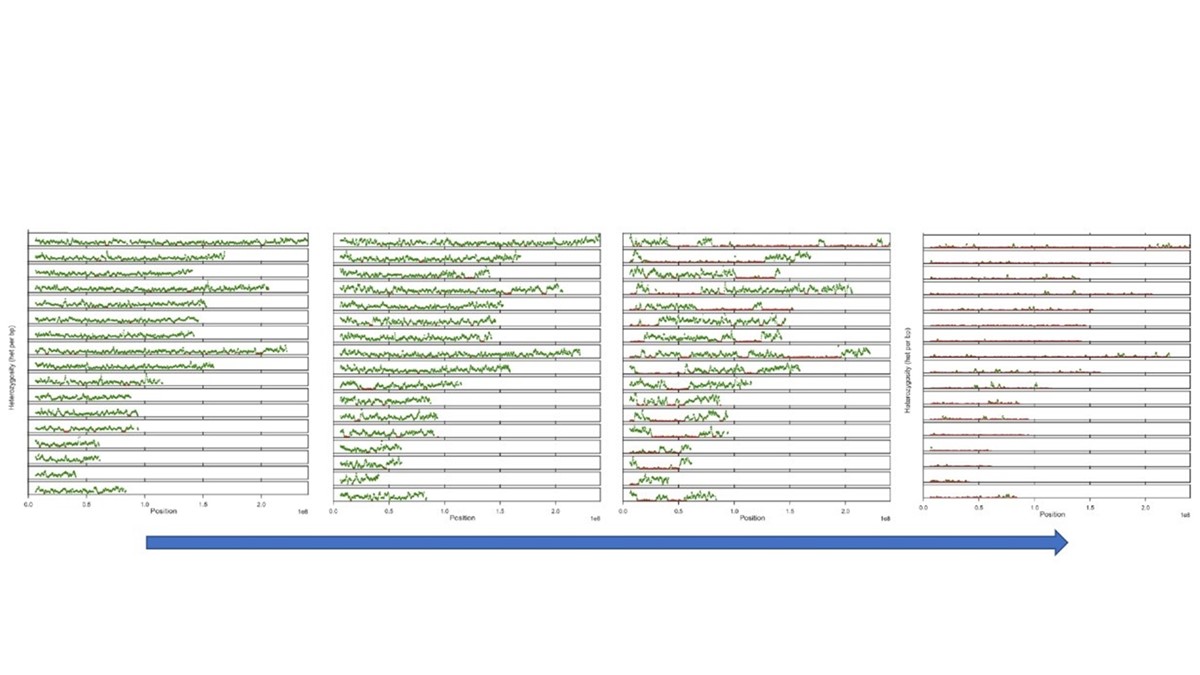
Organism populations are shaped by past events, some of which may have left long-lasting signatures in our genomes. Although numerous tools have been developed that enable such signatures to be detected using genomic datasets generated from contemporary materials, an alternative approach that is becoming increasingly feasible is to harness the power of population palaeogenomics - i.e. sequencing of ‘population’ scale datasets using ancient samples, chosen to span relevant locations and periods of interest. We have been exploring the potential of such methods in several systems, including domestic animals and plants, wild animals as well as humans in the context of one of the most notable catastrophes in recorded history - the second plague pandemic. In this talk I highlight the power of the population palaeogenomic approaches in producing new insights into such questions, using a range of our study systems.
Dr. Thomas Gilbert holds a PhD degree from the Zoology Department at Oxford University and subsequently, a post-doctoral fellowship at the Department of Ecology and Evolutionary Biology, University of Arizona. He became an Assistant Professor at the University of Copenhagen, where he is currently Professor of Palaeogenomics at the University of Copenhagen's Natural History Museum of Denmark, also leading the Gilbert group in the Centre for Geogenetics at Copenhagen University. He is also Adjunct Professor of Murdoch University (Perth, Australia). He is an evolutionary biologist and one of the most cited authors in the world, working on several topics, including: adaptive, comparative and speciation genomics; conservation and population genomics; domestication genomics; food genomics; metagenomics and metabarcoding; and paleogenomics.
[Host: BIODIV PhD Students]
Image credits: Thomas Gilbert