CIBIO-InBIO ADVANCED COURSE MULTIVARIATE STATISTICS FOR ECOLOGY AND EVOLUTION IN R
Many of the questions concerning the fields of ecology and evolution are better represented by multivariate datasets. The complex relationships observed across an ecological community, the association among different sets of phenotypic traits and the interaction between the phenotype and its environment are all fundamental questions in evolutionary ecology. The description and comprehension of complex traits is often better achieved through the quantification and analysis of multiple, frequently interdependent, phenotypic and ecological variables. The R-language for statistical computing has been increasingly used by evolutionary ecologists for statistical inference and hypothesis testing.
This course is directed towards PhD students interested in exploring the potential of R language for multivariate analyses in ecology and evolution. We will provide a general presentation of major statistical tools for multivariate analyses, including both exploratory and hypothesis-testing methods, and help the participants develop the necessary skills for implementing these tools using R. A basic previous knowledge of R is required and participants are responsible of guaranteeing the have the necessary basic R skills through a course of introduction to R or previous experience in working in this programming language.
COURSE INSTRUCTORS
Antigoni Kaliontzopoulou
Pedro Tarroso
COURSE OUTLINE
The course includes a morning (9:30-13:00) and an afternoon session (14:30-17:30).
During the morning session, we will discuss frequent questions in ecology and evolution and associate them to the statistical tools available for exploring them. The afternoon session will include a short demonstration of R code, followed by individual work by the participants. Participants are encouraged to bring their own data for analysis during the afternoon sessions. During the last day, each participant will give a short presentation of his/her study system, illustrating analyses explored during the practical sessions and/or describing how the tools the course may be useful in future analyses. This is also an opportunity for the students to get advice on data acquisition design and statistical treatment for on-going or planned studies.
DAY 1, June 29, 2015
1: Preliminary concepts: Why bother with multivariate analyses? Multivariate data spaces and metrics
2: General review of programming and the R environment: preparation for the practical sessions
DAY 2, June 30, 2015
3a: Inferential Methods I: Resampling (randomization, bootstrap, jackknife, Monte Carlo statistics)
3b: Inferential Methods II: Multivariate GLM (MANOVA, regression, MANCOVA)
DAY 3, July 1, 2015
4: Exploratory Methods: PCA, PCoA, MDS, clustering
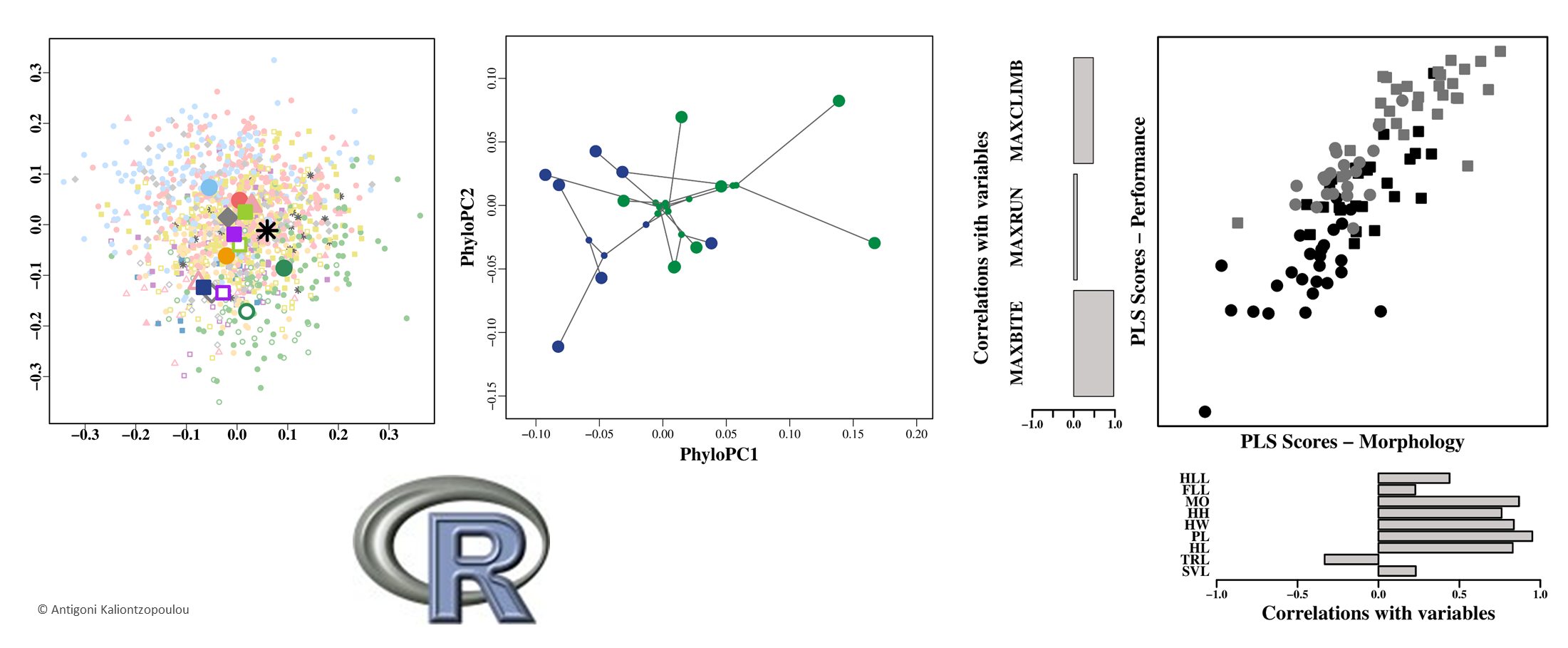
5: Association methods: PLS, CanCor, Mantel tests, CVA
DAY 4, July 2, 2015
6a: Methods for evolutionary and ecological non-independence I: phylogenetic comparative method
6b: Methods for evolutionary and ecological non-independence II: spatial considerations
DAY 5, July 3, 2015
Student presentations
All sessions will take place at Room 2.
INTENDED AUDIENCE
The course will be open to a maximum number of 20 participants.
Priority will be given to:
- 1st year PhD students attending the BIODIV Doctoral Program;
- Other BIODIV students;
- PhD students attending other courses;
- Post-doc researchers.
REGISTRATION
Registration deadline: June 15, 2015
Participation is free of charge for BIODIV students | Registration fee for other participants: € 100,00.
To register, please send an e-mail accompanied by your CV to Maria Sant’Ana at post.graduation@cibio.up.pt. Please refer your status (PhD student, MSc Student, Other) and the University to which you are affiliated.
![]()